
Metagenome.club
A metagenome is an environmental sample of genetic information, it can be made up of anything from algae in a pond to the microbiome in your stomach. My goal in this project was to work out a language for visualization and naming of a metagenome.
One of the primary challenges in metagenomics is that classical genetic barcoding methods and phylogenetic organization isn't an accurate representation of the cultural makeup of a metagenome. In a microbiome, genetic snippets are constantly being exchanged between species, making them far more dynamic than any current system is capable of representing.
The majority of visual representations of metagenomes take the form of phylogenetic trees, pie charts, or something that resembles a defragmenting hard drive. None of which really dealt with the dynamic aspect of a living culture made up colonies of similar yet different bacteria.
Given the limitations of time and the complicated nature of the topic I set out to simplify things for myself as much as possible.
I gave myself the goal of visualizing a metagenome over time and to come up with a system to give them common names.
I came up with a naming system based on the rules applied to hybridized daylilies, the choice was primarily because there are over 100,000 different hybridizations of daylilies each given a common name. And secondly because the names given to daylilies are amazing. daylily.org
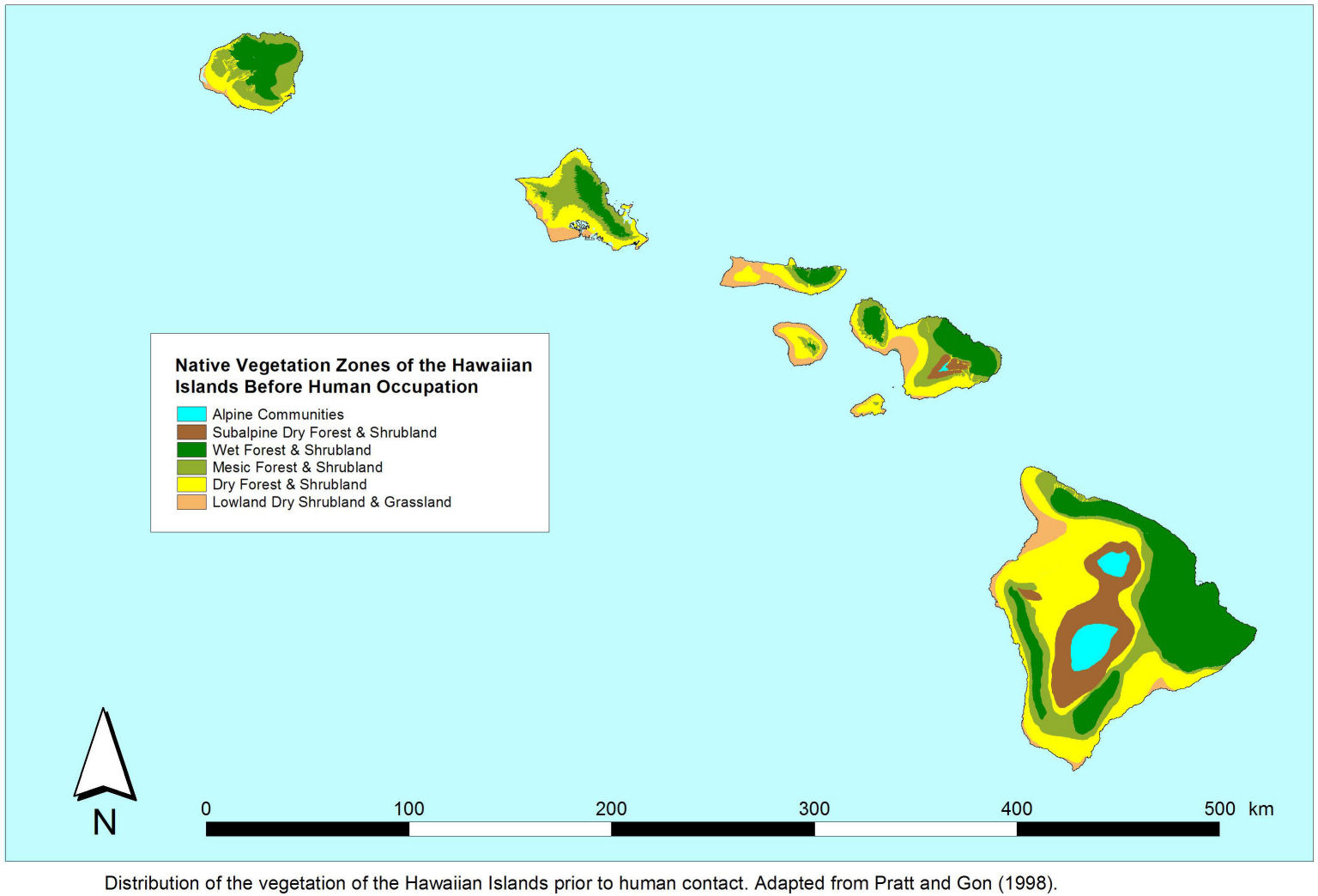
The visuals were inspired by the ecological maps of the hawaiian islands, however they ended up resembling bacteria more than islands in the end.
The visualization took form as a website called metagenome.club. The hope for the website (since it is still in development) is that it will operate like a panda cam, at any given moment you will be watching the same microbiome as anyone else watching, what is unique is that the organism itself will live on a server constantly evolving even when no one is watching.
At present the website generates a new name for every change in the state of the organism based on daylily names, in the future I propose that the viewer will be able to name the organism much like you would be able to name a star.
The organism is self documenting taking a photo of itself anytime someone is watching and a change occurs.
You can view the code for metagenome.club on github